Pathogen Evolutionary Genomics and Experimental Genetics
Microbial adaptability to the human host environment is crucial for disease causation. Pathogens like Plasmodium falciparum, Mycobacterium tuberculosis, and HIV-1, despite their differences in complexity, share the ability to replicate, persist, and transmit within hosts. The evolutionary and adaptive mechanisms enabling these capabilities also drive antimicrobial resistance, regulated at genomic and epigenomic levels.
The Pathogen Evolutionary Genomics and Experimental Genetics (PEvoGen) team employs cutting-edge functional and experimental genetics, including genome editing and multiple levels of gene-expression modulation, to explore how pathogen genetic variations impact biological functions. By integrating evolutionary genomics with “Omics” technologies and bioinformatics, PEvoGen provides novel insights into microbial pathogenicity, paving the way for innovative diagnostic and therapeutic solutions.
Understanding pathogen evolution through computational and experimental biology helps us anticipate and counter emerging infectious threats, particularly antimicrobial resistance. This research aligns with the principles of the One Health concept and supports the Sustainable Development Goals (SDGs).
Team Members


Nuno S. Osório

Ana Santos-Pereira

Claudia Fançony

Vitória da Cunha Baptista

Beatriz Velosa da Fonseca

Bruno José Ferreira Freitas
Domingos Oliveira

Filipa Tavares

Flávia Figueira

Maria Eduarda Pereira Mascarenhas

Maria Pereira

Matheus Tavares
Nuno Alves
Plácida Maholela

Ruth Victoria Esho

Sarah Brena Rosa

Talita Nicolau

Filipe Barbosa de Vasconcelos

Margarida Gonçalves

Tiago Gonçalves
Team Members

Maria Isabel Veiga
Principal Investigator
Team coordinator

Nuno S. Osório
Senior Researcher

Ana Santos-Pereira
Postdoctoral Researcher

Claudia Fançony
Postdoctoral Researcher

Vera Cristina Aguieiras Triunfante
Affiliated Researcher

Carlos Magalhães
PhD Student

Maria Pereira
PhD Student

Pedro Araújo
PhD Student

Vitória da Cunha Baptista
PhD Student

Beatriz Velosa da Fonseca
MSc Student

Bruno José Ferreira Freitas
MSc Student

Claudia Alejandra Cabrera Federo
MSc Student

Filipa Tavares
MSc Student

Inês Gonçalves Geraldes
MSc Student

Mariagiulia Conte
MSc Student

Ruth Victoria Esho
Affiliated Researcher
Projects
- Ana Santos-Pereira
- There is still no cure for HIV infection and the extensive viral genetic diversity represents the one of the biggest obstacles to develop an effective therapy…
- Maria Isabel Veiga
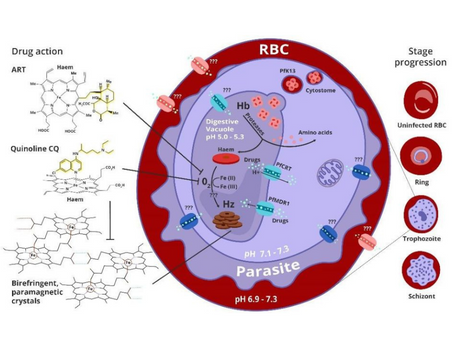
- The clinical manifestation of malaria starts when parasites invade human red blood cells (RBCs) and feed on haemoglobin, releasing a toxic by-product, polymerized by the parasite into the inert crystal hemozoin..
- Maria Isabel Veiga
- Hampering malaria control and elimination efforts is the resilient capacity of the parasite Plasmodium falciparum to develop resistance, including resistance to the presently…
- Vitória Baptista
- This project aims to unravel the molecular mechanisms behind hemozoin formation in Plasmodium falciparum, a vital process for parasite survival and a key target of antimalarial drugs.
- Maria Isabel Veiga
- This project aims to develop a machine learning model to predict malaria treatment failure at the patient level, enabling more personalized and effective care…
- Nuno S. Osório
- Tuberculosis (TB) is one of the oldest and deadliest human infectious disease. Only innovative tools will allow realistic approximations to TB eradication…
- Nuno S. Osório
- The extremely high human-to-human transmissibility of SARS-CoV-2 allows the generation of thousands of mutations within its genome fueling the evolution and selection of several viral lineages with different characteristics…
- Claudia Fançony
- There is still no cure for HIV infection and the extensive viral genetic diversity represents the one of the biggest obstacles to develop an effective therapy…
Success Story
Selected Research Outputs
WO2021191882 ANTIMALARIAL AGENT, METHODS AND USES THEREOF (wipo.int)
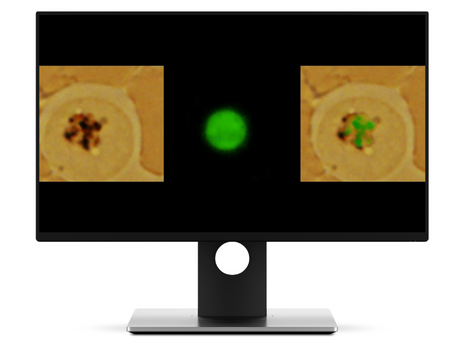
WO2022070084 AUTOMATIC DEVICE FOR NON-INVASIVE MALARIA DIAGNOSIS THROUGH OPTICAL REFLECTANCE TECHNIQUES, METHODS AND USES THEREOF (wipo.int)
Frontiers | Plasmodium falciparum Drug Resistance Genes pfmdr1 and pfcrt In Vivo Co-Expression During Artemether-Lumefantrine Therapy (frontiersin.org)
The Future in Sensing Technologies for Malaria Surveillance: A Review of Hemozoin-Based Diagnosis | ACS Sensors
PEvoGen team main achievements:
– Contributed to the knowledge of the mechanisms of resistance to antimalarial treatments in the malaria parasite Plasmodium falciparum
– Improved antimalarial drug discovery by identifying new, essential, druggable targets and test compounds that may be advanced into drug candidates suitable for preclinical development and subsequent clinical testing in humans.
– Explored better malaria diagnosis, using differential physical properties of the malaria parasite regarding the hemozoin biological pathway.
– Highlighted relevant factors contributing to the colonization of human populations by viruses of different HIV-1 subtypes.
– Endeavored the challenge of investigating Mycobacterium tuberculosis mutations accumulated during the transmission between individuals infected in the same transmission chain.
– Advanced the study of phylogeny, transmission and dissemination of HIV-1, M. tuberculosis and more recently SARS-CoV-2 through Bioinformatics.
Former lab members
• Carla Sofia Calçada, PhD (Scientific Researcher, Toronto, Canada)
• Deisy Rocha, PhD (Postdoctoral Researcher, IPBS-Toulouse, France)
• Diana Granja, MSc (Bioengineer, Anocca biotech company, Sweden)
• Francisco Araujo, MD, MSc (Project leader, Autonomous University of Santo Domingo, Dominican Republic)
• Geniss Nhapulo, MD
• Isaac Sanchéz, MSc (Technical Coordinator at the National Reference Public Health Laboratory, Dominican Republic)
• Leyre Lau (PhD student, Karolinska Institute, Sweden)
• Miguel João Ferreira da Silva, PhD (Postdoctoral Researcher, Masaryk University, Czechia)
Evolutionary dynamics of HIV-1 subtype C in Brazil | Scientific Reports (nature.com)
Biomedicines | Free Full-Text | OmniSARS2: A Highly Sensitive and Specific RT-qPCR-Based COVID-19 Diagnostic Method Designed to Withstand SARS-CoV-2 Lineage Evolution (mdpi.com)
Plasmodium falciparum transcription in different clinical presentations of malaria associates with circulation time of infected erythrocytes | Nature Communications
Implication of SARS-CoV-2 evolution in the sensitivity of RT-qPCR diagnostic assays - The Lancet Infectious Diseases
Expansion of a Specific Plasmodium falciparum PfMDR1 Haplotype in Southeast Asia with Increased Substrate Transport | mBio
Mycobacterium tuberculosis associated with severe tuberculosis evades cytosolic surveillance systems and modulates IL-1β production | Nature Communications



Contact us
Phone: +351 253 604 967
Fax: +351 253 604 809
Email: icvs.sec@med.uminho.pt
Address
Life and Health Sciences
Research Institute (ICVS)
School of Medicine,
University of Minho,
Campus de Gualtar
4710-057 Braga
Portugal

Copyright ©2025 ICVS. All Rights Reserved. Developed by TCIT



Copyright ©2025 ICVS. All Rights Reserved. Developed by TCIT
Address
Life and Health Sciences
Research Institute (ICVS)
School of Medicine,
University of Minho,
Campus de Gualtar
4710-057 Braga
Portugal



Copyright ©2025 ICVS. All Rights Reserved
Address
Life and Health Sciences
Research Institute (ICVS)
School of Medicine,
University of Minho,
Campus de Gualtar
4710-057 Braga
Portugal